Validation report: extend_v1¶
Model: extend_v1
Samples Generated: 500
Generated On: 2026-01-05 00:28
Generation Quality Metrics¶
Uniqueness |
Novelty |
SELFIES fidelity |
|---|---|---|
0.9960 |
0.6426 |
0.0152 |
Descriptor Statistics¶
Descriptor |
Average |
Minimum |
Maximum |
|---|---|---|---|
MW |
246.2045 |
97.1600 |
496.0100 |
LogP |
1.9321 |
-2.2100 |
7.6400 |
SA_Score |
3.5984 |
1.5400 |
6.9800 |
QED |
0.6220 |
0.1950 |
0.9430 |
Fsp3 |
0.5701 |
0.0000 |
1.0000 |
RotatableBonds |
3.0820 |
0.0000 |
12.0000 |
RingCount |
2.5800 |
0.0000 |
9.0000 |
TPSA |
47.8160 |
0.0000 |
138.8300 |
RadicalElectrons |
0.0120 |
0.0000 |
2.0000 |
Descriptor Distributions¶
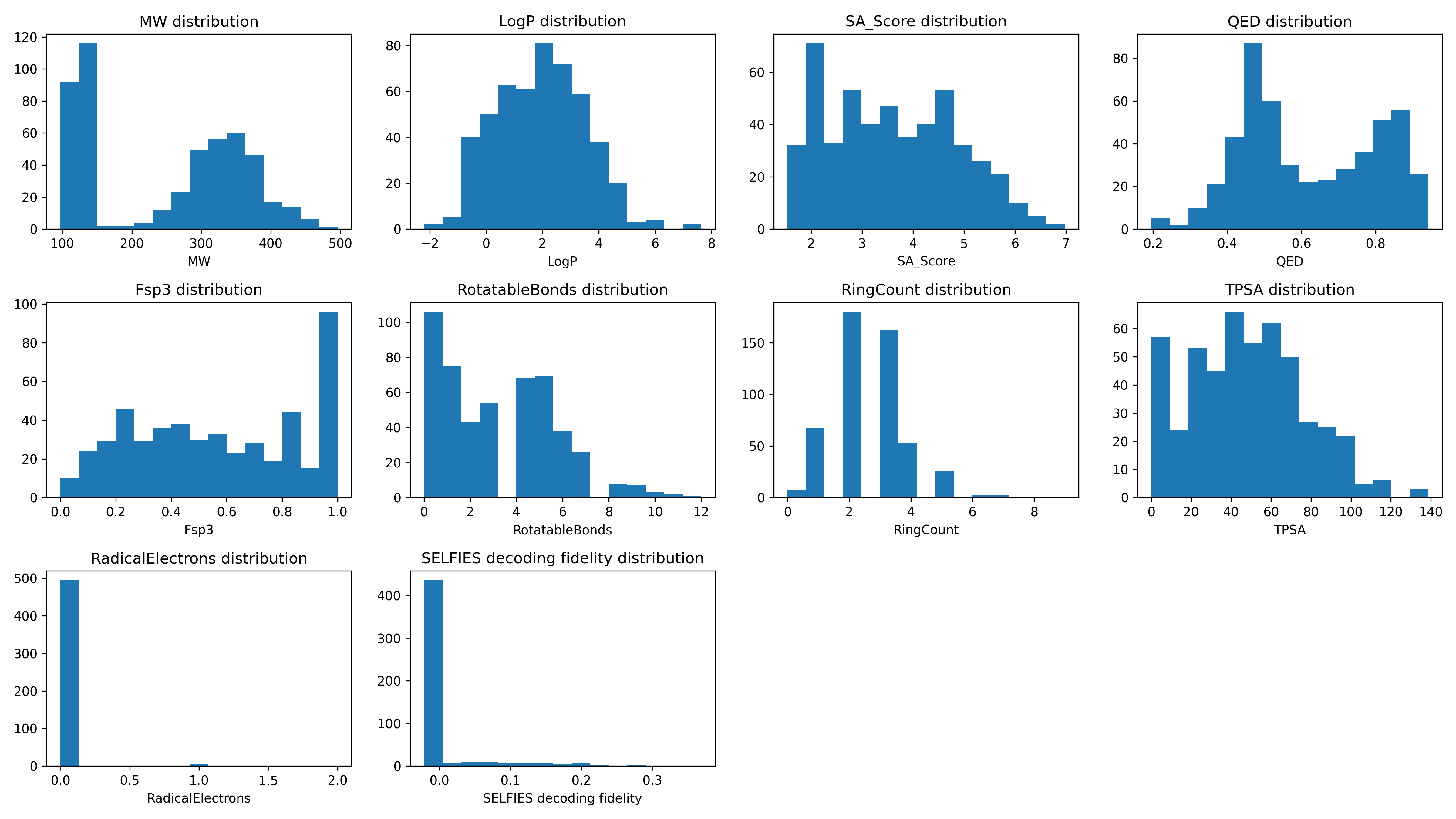
Development Notes¶
Model was trained on SMILES patterns encoded into SELFIES. Training data was obtained from QM9 and ZINC datasets. Training was stopped when a plateau in loss was observed even with a reduced learning rate of 1e-5.
PyTorch dataset was defined as follows:
class ChempleterDataset(Dataset):
"""
PyTorch Dataset for SELFIES molecular representations.
:param selfies_file: Path to CSV file containing SELFIES strings in a "selfies" column.
:type selfies_file: str
:param stoi_file: Path to JSON file mapping SELFIES symbols to integer tokens.
:type stoi_file: str
:returns: Integer tensor representation of tokenized molecule with dtype=torch.long.
:rtype: torch.Tensor
"""
def __init__(self, selfies_file, stoi_file):
super().__init__()
selfies_dataframe = pd.read_csv(selfies_file)
self.data = selfies_dataframe["selfies"].to_list()
with open(stoi_file) as f:
self.selfies_to_integer = json.load(f)
def __len__(self):
return len(self.data)
def __getitem__(self, index):
molecule = self.data[index]
symbols_molecule = ["[START]"] + list(sf.split_selfies(molecule)) + ["[END]"]
integer_molecule = [
self.selfies_to_integer[symbol] for symbol in symbols_molecule
]
return torch.tensor(integer_molecule, dtype=torch.long)
